leukemia-research.de
Research Group, Pediatric Hematology and Oncology
Headed by Prof. Dr. Jan-Henning Klusmann and Prof. Dirk Heckl, we aim to understand the molecular basis underlying the development of myeloid leukemia. In order to achieve this goal and model this disease, we use state-of-the-art techniques including a variety of CRISPR-based techniques and next generation sequencing.
RESEARCH BLOG
Learn about early results as they happen.
Leukemia research on Television
The latest episode of a regional news outlet on German television network SAT.1 featured Prof.Klusmann and Prof.Heckl. The interview gives insights on possible applications of micro RNAs in cancer therapies.
Luca wins Abstract Award at ASH 2023
Luca was awarded with the abstract award and presents his work at this years annual meeting of the American Society of Hematology in San Diego.
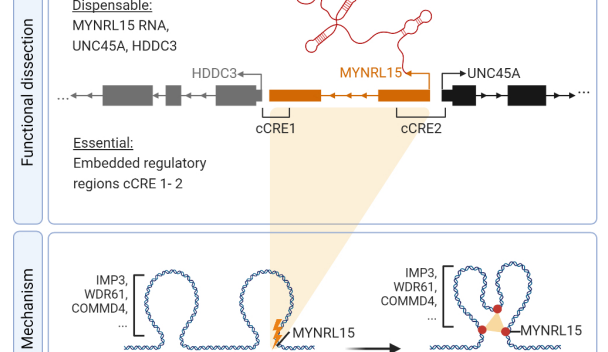
Myeloid leukemia vulnerabilities embedded in long non-coding RNA locus MYNRL15
Publication alert: The identification of long non-coding RNA locus MYNRL15 shows novel therapeutic potential in AMLs with various genetic backgrounds.
RESEARCH PROJECTS
Explore our daily work in science.
Long non-coding RNAs
In the post-genomic era, the ongoing discovery of transcriptional products from non-coding regions of the genome continues to raise fundamental questions of biology. Long non-coding RNAs are a versatile class of transcripts, which add a new layer of complexity to the regulatory networks behind normal and malignant hematopoiesis.
MicroRNAs
MicroRNAs are short, non-coding, master post-transcriptional regulators of gene expression. They have been established as key players in developmental and physiological processes, as well as in pathological ones such as cancer. We have long been interested in the roles of microRNAs during both leukemic transformation and normal blood formation.
GATA1s in Down syndrome AML
Children with Down syndrome (trisomy 21) have a 20-fold greater risk of developing myeloid leukemia. Of neonates with Down syndrome, 10% are diagnosed with transient leukemia, of which 30% develop full-blown leukemia within three years. We are investigating the additional genetic hits that co-operate with GATA1s to drive this progression.
CRISPR-Cas9 methods to model AML
The discovery and application of CRISPR-Cas9 has enhanced genome editing, overcoming a number of the limitations seen with previous genome editing tools. The system is efficient and simple, and acts at the endogenous locus on cellular DNA – both a major advance and advantage, with respect to achieving more physiological conditions.
Battling leukemia daily
Exploring the basis of leukemia with functional genomics.
The first step towards developing specific therapies – that eradicate leukemia cells but spare normal cells of the human body – is to understand how leukemia evolves. The key purpose of our daily work is to link genetic perturbations to biological processes involved in leukemia. This simple principle is central to our research efforts and is true for reverse-genetic (genotype-to-phenotype) as well as forward-genetic approaches (phenotype-to-genotype). In our work, molecular genetics and the struggle to cure leukemia thus meet on a daily basis.
By applying genome editing, we elucidate the molecular processes induced by numerous known mutations that we and others identified in leukemia genomes. This has enabled new understanding of leukemic transformation in children with Down syndrome, and of Polycomb Repressive Complex 2 and chromosomal rearrangements in pediatric leukemia. Beyond genetic mutations, we mapped transcriptional landscapes of leukemia and normal hematopoiesis, providing further insights into the cellular networks hijacked by this disease.
Refining the genotype-phenotype correlations in leukemia permits us to next uncover weaknesses and markers that improve patient survival. Towards this goal, we apply forward-genetic screens based on gene inactivation and targeted perturbation of gene expression (CRISPRi/a), via the CRISPR-Cas9 system. With the integration of known drug response profiles, these studies will help us develop new treatment strategies – a point we have proven by introducing histone deacetylase inhibitors as an effective class of drugs for Down syndrome leukemia.
About our scientific work
The overarching aim of our research is to develop novel, cancer-specific, patient-orientated treatment options for children with acute leukemia.
Among the different types of leukemia, our research primarily focuses on acute myeloid leukemias, a subtype in which treatment outcomes have not substantially improved for pediatric patients for decades. We are particularly interested in children with Down syndrome, who are predisposed towards developing myeloid leukemia, and who present with increased sensitivity to treatment-related toxicities. Over the years, we have contributed to identifying coding and non-coding genes on chromosome 21 that are involved in disease pathogenesis. We also uncovered a complex transcriptional network that is deregulated during malignant transformation, at the center of which is GATA1s – a crucial factor for Down syndrome myeloid leukemia development. Utilizing advanced molecular biology approaches, we aim to delineate the interaction partners of GATA1s, the oncogenic network on chromosome 21, and co-occurring mutations during the transformation into Down syndrome myeloid leukemia.
Another of our central interests is the role of non-coding RNAs in acute myeloid leukemia. Considered to be cellular “junk” for decades, insights into the transcriptome have since revealed their key functions as regulators of gene expression. With our non-coding RNA atlas of the human blood system covering the entire hematopoietic hierarchy as well as pediatric acute myeloid leukemia samples, we have built a foundation from which to decipher pathologic mechanisms guided by non-coding RNAs.