Project Description
Long non-coding RNAs in (de)regulated hematopoiesis
In the post-genomic era, the ongoing discovery of transcriptional products from non-protein-coding areas of the genome continues to raise fundamental questions of biology. Only 2% of the human genome encodes protein, but 70-90% is transcribed, resulting in an enormous number and diversity of non-coding transcripts. In recent years, new classes of non-coding RNAs were ascribed with regulatory functions in the cell, including microRNAs (miRNAs) and long non-coding RNAs (lncRNAs).
While miRNAs have been established for a while, our lab is also interested in lncRNAs, which are only emerging as novel regulators of gene expression. Loosely defined as non-coding transcripts longer than 200 nucleotides, lncRNAs can be sense, antisense, overlapping or intergenic to protein-coding loci, and are versatile molecules capable of binding DNA, RNA or protein. Large-scale studies have revealed their developmental stage- and cellular context-specific expression patterns; however, the functional work still lags behind in silico discoveries.
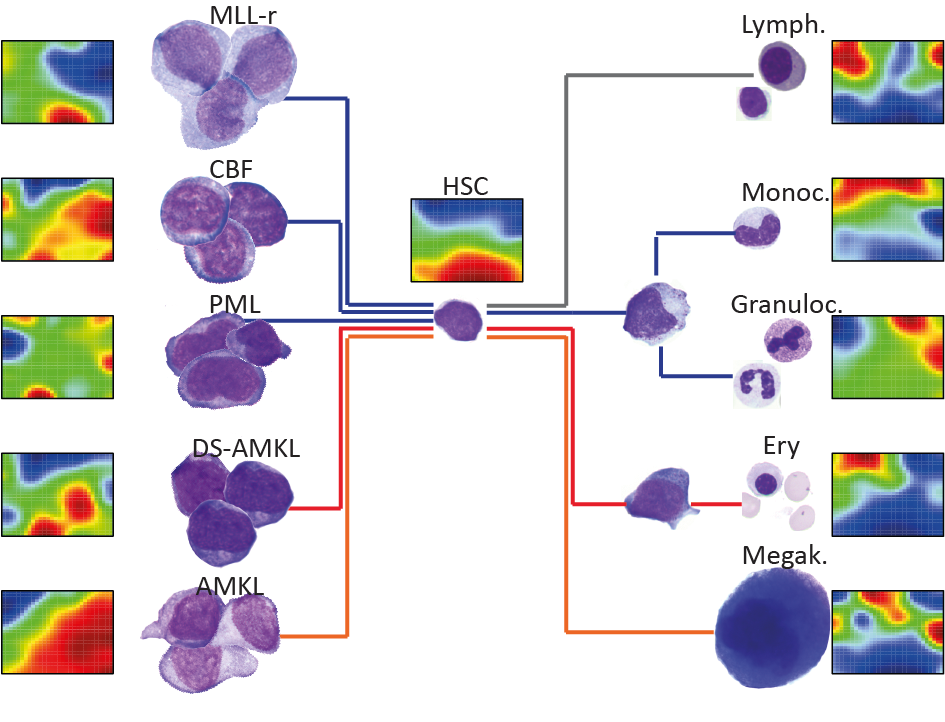
To address this knowledge gap in the context of hematopoiesis, we built a non-coding expression atlas for the human blood system, with profiles of hematopoietic stem cells, differentiated blood lineages, as well as pediatric AML samples. We were able to identify lineage and leukemia-specific lncRNAs, and further, stem cell and progenitor lncRNA signatures that are upregulated in AML. Based on these findings, we are working to elucidate the functions of lncRNAs during normal and malignant hematopoiesis.
The stem cell-specific long noncoding RNA HOX10-AS in the pathogenesis of KMT2A-rearranged leukemia.
Al-Kershi S, Bhayadia R, Ng M, Verboon L, Emmrich S, Gack L, Schwarzer A, Strowig T, Heckl D, Klusmann JH.
Blood Adv. 2019 Dec 23;3(24):4252. doi: 10.1182/bloodadvances.2019032029
The regulatory roles of long noncoding RNAs in acute myeloid leukemia.
Ng M, Heckl D, Klusmann JH.
Front Oncol. 2019 Jul 9;9:570. doi: 10.3389/fonc.2019.00570.
The non-coding RNA landscape of human hematopoiesis and leukemia.
Schwarzer A, Emmrich S, Schmidt F, Beck D, Ng M, Reimer C, Adams FF, Grasediek S, Witte D, Käbler S, Wong JWH, Shah A, Huang Y, Jammal R, Maroz A, Jongen-Lavrencic M, Schambach A, Kuchenbauer F, Pimanda JE, Reinhardt D, Heckl D, Klusmann JH.
Nat Comm. 2017 Aug 9;8(1):218. doi: 10.1038/s41467-017-00212-4.